7 Reasons Why Power BI is a Game-Changer for Organizations
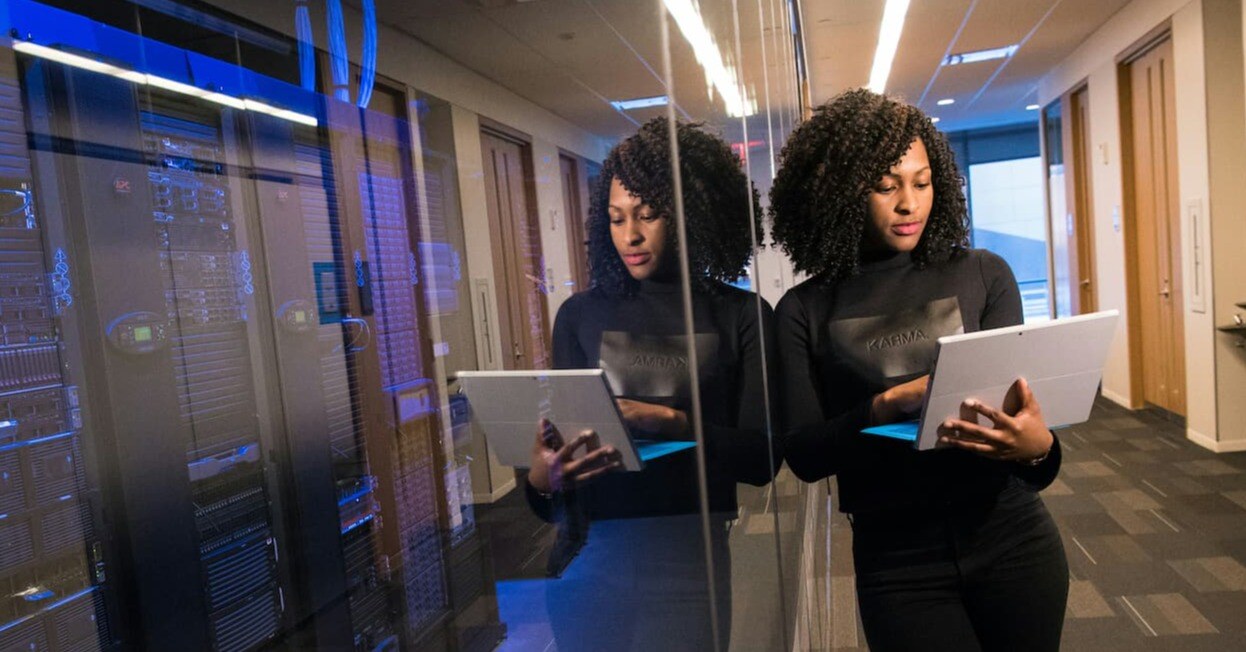
The amount of data we have accessible has never been greater. Data on our customers, the markets we serve, and data from nearly every device we use. The challenge lies in whether we know how to use this data to our advantage.